 
Hydrogen bonding patterns in secondary structures may be significantly distorted, which makes automatic determination of secondary structure difficult. Neutron scattering measurements have directly connected the spectral feature at ~1 THz to collective motions of the secondary structure of beta-barrel protein GFP. Low frequency collective vibrations are thought to be sensitive to local rigidity within proteins, revealing beta structures to be generically more rigid than alpha or disordered proteins. However, these preferences are not strong enough to produce a reliable method of predicting secondary structure from sequence alone. Amino acids that prefer to adopt helical conformations in proteins include methionine, alanine, leucine, glutamate and lysine ("MALEK" in amino-acid 1-letter codes) by contrast, the large aromatic residues ( tryptophan, tyrosine and phenylalanine) and C β-branched amino acids ( isoleucine, valine, and threonine) prefer to adopt β-strand conformations. Proline and glycine are sometimes known as "helix breakers" because they disrupt the regularity of the α helical backbone conformation however, both have unusual conformational abilities and are commonly found in turns. The random coil is not a true secondary structure, but is the class of conformations that indicate an absence of regular secondary structure.Īmino acids vary in their ability to form the various secondary structure elements. Tight turns and loose, flexible loops link the more "regular" secondary structure elements. Other extended structures such as the polyproline helix and alpha sheet are rare in native state proteins but are often hypothesized as important protein folding intermediates. Other helices, such as the 3 10 helix and π helix, are calculated to have energetically favorable hydrogen-bonding patterns but are rarely observed in natural proteins except at the ends of α helices due to unfavorable backbone packing in the center of the helix. The most common secondary structures are alpha helices and beta sheets. Cartoon above, atoms below with nitrogen in blue, oxygen in red ( PDB: 1AXC) The image above contains clickable links Interactive diagram of hydrogen bonds in protein secondary structure. Types Structural features of the three major forms of protein helices Geometry attribute Other types of biopolymers such as nucleic acids also possess characteristic secondary structures. The concept of secondary structure was first introduced by Kaj Ulrik Linderstrøm-Lang at Stanford in 1952. Secondary structure may alternatively be defined based on the regular pattern of backbone dihedral angles in a particular region of the Ramachandran plot regardless of whether it has the correct hydrogen bonds. Secondary structure is formally defined by the pattern of hydrogen bonds between the amino hydrogen and carboxyl oxygen atoms in the peptide backbone. Secondary structure elements typically spontaneously form as an intermediate before the protein folds into its three dimensional tertiary structure. The two most common secondary structural elements are alpha helices and beta sheets, though beta turns and omega loops occur as well.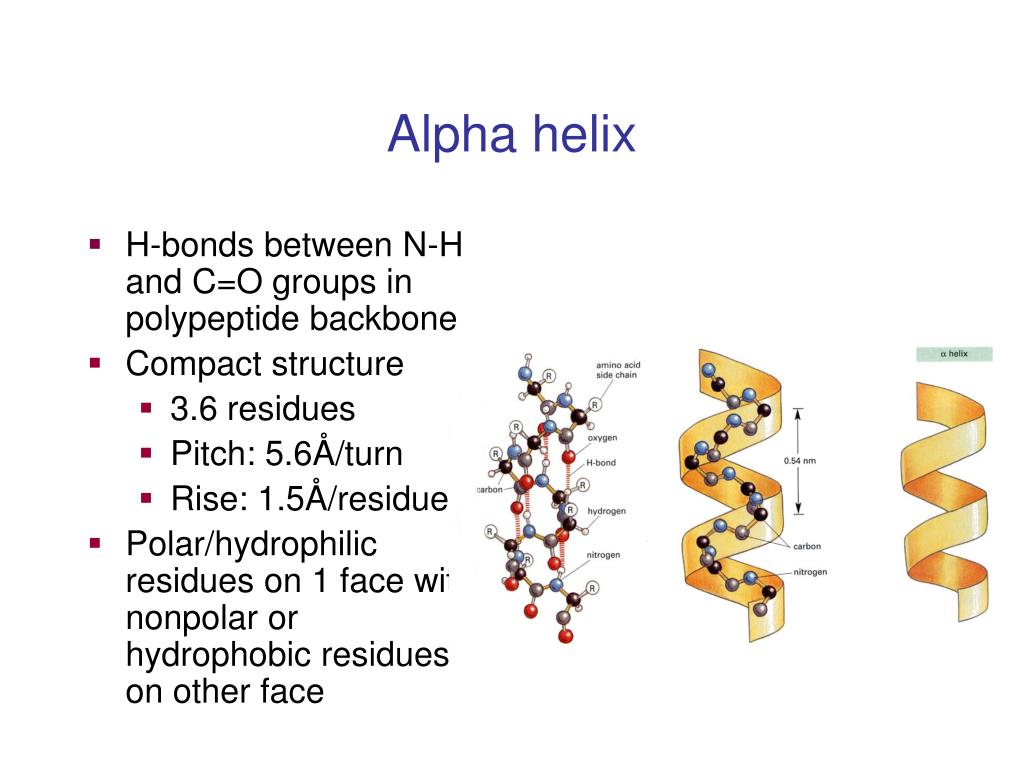 
Protein secondary structure is the local spatial conformation of the polypeptide backbone excluding the side chains. 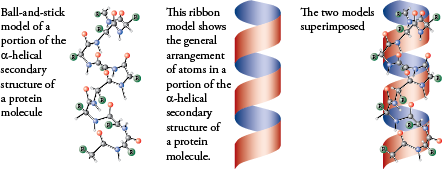
The image above contains clickable links This diagram (which is interactive) of protein structure uses PCNA as an example. For the article about secondary structure in nucleic acid, see Nucleic acid secondary structure. This article is about secondary structure in protein.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
